Research Article
Distribution, Antibiotic Sensitivity and Biofilm Formation of Nosocomial Pathogens Isolated in Male and Female Surgical Ward Environments from a Tertiary Hospital in Ado-Ekiti, Ekiti State
1 Department of Medical Laboratory Science, Faculty of Basic Medical Science, Ekiti State University, Ado-Ekiti, Nigeria.
2 Department of Microbiology, Faculty of Science, Ekiti State University, Ado-Ekiti, Nigeria.
3 Department of Molecular and Medical Biotechnology, University of Verona, Italy.
4 Department of Biochemistry, Faculty of Natural and Applied Sciences, Lead City University, Ibadan, Oyo State, Nigeria.
*Corresponding Author: F.A. Adewumi, Department of Medical Laboratory Science, Faculty of Basic Medical Science, Ekiti State University, Ado-Ekiti, Nigeria.
Citation: Adewumi F. A., Ipinlaye J. O., Owoyemi A. C., Alonge F. B., Airaodion A. I., et al. (2026). Distribution, Antibiotic Sensitivity and Biofilm Formation of Nosocomial Pathogens Isolated in Male and Female Surgical Ward Environments from a Tertiary Hospital in Ado-Ekiti, Ekiti State, Journal of Clinical Infectious Diseases and Reports, BioRes Scientia Publishers. 2(1):1-8. DOI: 10.59657/jcidr.brs.26.007
Copyright: © 2026 F. A. Adewumi, this is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Received: January 12, 2026 | Accepted: January 26, 2026 | Published: February 03, 2026
Abstract
Background: Nosocomial infections are hospital-acquired infections that occur during a stay of 48-72 hours. It results from transmission during hospital care through the use of contaminated devices or healthcare workers. Worldwide, 5-10% of patients develop Healthcare-Associated Infections (HAIs), which have been a global threat to human health.
Aims: This study aims to track the prevalence of antibiotic-resistant and biofilm-forming bacteria on environmental surfaces in the tertiary hospital.
Methodology: A total of 33 environmental samples were analyzed, comprising 12 tables, 8 drawers, 6 chairs, 6 irons, and a bed. Isolates were cultured on MacConkey agar and Sheep blood agar using the streaking method. Isolates identification was carried out by performing a series of biochemical tests and their cultural morphology on MacConkey, Cetrimide, Mannitol salt and Eosin methylene blue (EMB) agar. Polymerase chain reaction was employed for the confirmatory identification of the isolated recovered. Antibiotic susceptibility was done using the disc diffusion method. Phenotypic Carbapenem test was carried out on isolates to identify multi-drug Antibiotic resistant isolates (MDR), while biofilm formation test was quantified using crystal violet assay.
Result: The result of this study revealed that out of 33 environmental samples analyzed, Escherichia coli had the highest frequency, 10 (30.3%), followed by Pseudomonas aeruginosa 8 (24%), Proteus mirabilis, 8 (24%) and Klebsiella pneumonia, 5 (15.5%). Staphylococcus aureus had the lowest frequency of 2 (6.06%). According to the antibiotic susceptibility test, fifteen out of thirty-three isolates (4.54%) have multiple resistance to ≥7 antibiotics. Also, thirty out of thirty-three isolates were resistant to AUG (Augmentin), CTX (Ceftriaxone), and IMP (Imipenem). The highest resistance to meropenem 8 (22.2%), ertapenem 7 (19.4%), and imipenem 5 (13.8%) is seen in table samples, as indicated in the carbapenem results. The biofilm results demonstrated that Escherichia coli and Klebsiella pneumonia have a strong biofilm-forming potential. It was observed that isolates from tables were positive in the biofilm formation test.
Conclusion: This study identified clinically significant nosocomial pathogens such as Escherichia coli, Staphylococcus aureus, Klebsiella pneumoniae, Pseudomonas aeruginosa and Proteus mirabilis from environmental surfaces within male and female surgical wards of a tertiary healthcare facility. The detection of multidrug-resistant and biofilm-forming bacteria highlights an urgent need for strengthened infection prevention and control measures. The high prevalence of resistance underscores the necessity for robust antimicrobial stewardship programs and improved environmental hygiene practices.
Keywords: nosocomial; antibiotics; multidrug resistance; carbapenem; biofilms
Introduction
Nosocomial infections, also referred to as hospital-acquired infections (HAIs), are infections that develop during hospitalisation and were neither present nor incubating at the time of a patient’s admission. Conventionally, these infections become clinically evident 72 hours or more after admission [1]. HAIs occur across diverse healthcare settings and affect both patients and healthcare personnel. They are caused by a wide range of microorganisms, including bacteria, fungi, viruses, and parasites, with bacteria accounting for the majority of reported cases [2]. Common bacterial agents implicated in nosocomial infections include Streptococcus species, Acinetobacter species, Bacillus cereus, Enterococcus species, Pseudomonas aeruginosa, coagulase-negative staphylococci, Staphylococcus aureus, Legionella species, and members of the Enterobacteriaceae family.
Globally, nosocomial infections contribute significantly to patient morbidity and mortality, while also imposing substantial economic burdens on healthcare systems. Similar patterns have been documented in Nigeria, where HAIs remain a major cause of prolonged hospital stay, increased healthcare costs, and avoidable deaths among hospitalised patients [3]. The most frequently reported HAIs include hospital-acquired urinary tract infections, hospital-acquired pneumonia, upper respiratory tract infections, skin and soft tissue infections, surgical site infections, ventilator-associated pneumonia, and Clostridioides difficile infections, with bacterial pathogens responsible for approximately 90% of cases [4]. Non-compliance with established infection prevention and control guidelines has been identified as a key driver of HAI transmission within hospital environments [5].
A critical factor that enhances bacterial persistence in hospital settings is the ability to form biofilms on both living and non-living surfaces [6]. Biofilms are structured microbial communities encased within a self-produced extracellular polymeric substance (EPS) matrix. This matrix is composed of polysaccharides, proteins, lipids, extracellular DNA and RNA, and other macromolecules that collectively provide mechanical stability and protection to the microbial cells [7]. Biofilm formation represents an important virulence trait, as it markedly increases bacterial tolerance to antibiotics and disinfectants. Biofilm-associated infections are therefore difficult to eradicate and frequently result in chronic or recurrent disease. It is estimated that between 65% and 80% of bacterial infections in humans involve biofilm-forming organisms [8,9].
In hospital wards, nosocomial pathogens pose a serious threat due to their ease of dissemination, high levels of antimicrobial resistance, and strong biofilm-forming capacity. At Ekiti State University Teaching Hospital (EKSUTH), limited information is available regarding the distribution, antimicrobial susceptibility patterns, and biofilm-forming potential of nosocomial bacteria present on environmental surfaces in male and female surgical wards. This knowledge gap hampers effective infection control strategies and may contribute to increased patient morbidity and mortality [10,11]. The present study was therefore designed to investigate bacterial contamination of hospital surfaces, assess antibiotic susceptibility profiles, and evaluate biofilm formation among isolates, with the overarching goal of identifying effective therapeutic options and improving clinical outcomes.
Materials and Methods
Sample Collection
This study was carried out in the male and female surgical wards of a tertiary healthcare facility located in Ado-Ekiti, Ekiti State, Nigeria. A total of 33 environmental swab samples were collected over a one-month period. The sampled surfaces included 12 tables, 8 drawers, 6 chairs, 6 iron surfaces, and 1 bedding. Samples were obtained aseptically using sterile swab sticks, appropriately labelled, transported under cold conditions, and processed promptly in the microbiology laboratory.
Microbiological Analysis
Bacterial isolation and identification were performed using standard microbiological techniques. Isolates were characterised based on colony morphology, Gram reaction, and conventional biochemical tests following established laboratory protocols [12].
Antibiotic Susceptibility Testing
Antibiotic susceptibility testing was conducted using the agar disc diffusion method in accordance with Clinical and Laboratory Standards Institute (CLSI) guidelines [13]. Fourteen antibiotics were tested: cefotaxime (CTX, 25 μg), imipenem (IMP, 10 μg), ofloxacin (OFX, 5 μg), gentamicin (GN, 10 μg), nalidixic acid (NA, 30 μg), nitrofurantoin (NF, 300 μg), cefuroxime (CXM, 30 μg), ceftriaxone–sulbactam (CRO, 45 μg), ampliclox (ACX, 10 μg), cefixime (ZEM, 5 μg), levofloxacin (LBC, 5 μg), ciprofloxacin (CIP, 5 μg), erythromycin (ERY, 5 μg), and azithromycin (AZN, 15 μg).
Pure bacterial colonies were inoculated into nutrient broth and incubated at 35 °C until turbidity equivalent to the 0.5 McFarland standard was achieved. Mueller–Hinton agar plates were inoculated and antibiotic discs applied aseptically. Plates were incubated at 37 °C for 24 hours, after which inhibition zone diameters were measured and interpreted as susceptible, intermediate, or resistant using CLSI breakpoints [13].
Phenotypic Detection of Carbapenem Resistance
Carbapenem resistance was evaluated phenotypically using the disk diffusion method following CLSI recommendations [13]. Standardised bacterial suspensions were inoculated onto Mueller–Hinton agar plates, and ertapenem (ETP, 10 μg), imipenem (IMP, 10 μg), and meropenem (MEM, 10 μg) discs were applied. Plates were incubated aerobically at 35–37 °C for 16–18 hours, and inhibition zones were measured and interpreted accordingly.
Molecular Identification of Bacterial Isolates by PCR
DNA Extraction
Genomic DNA was extracted from overnight cultures using the boiling method. Briefly, bacterial colonies were suspended in sterile distilled water, heat-lysed at 95 °C for 10 minutes, rapidly cooled, centrifuged, and the supernatant containing DNA was used as PCR template.
PCR Amplification and Gel Electrophoresis
Species-specific primers were used to detect Escherichia coli, Pseudomonas aeruginosa, Proteus mirabilis, Klebsiella pneumoniae, and the mecA gene. Amplification products were resolved on 1.5% agarose gels stained with ethidium bromide and visualised under ultraviolet illumination using a 100 bp DNA ladder as a molecular marker.
Table: Species-Specific Primers Used
| Organism | Target Gene | Primer Sequence (5′–3′) | Amplicon Size (bp) | Reference |
| Escherichia coli | uidA | F: TGGTAATTACCGACGAAAACGGC | ||
| R: ACGCGTGGTTACAGTCTTGCG | 147 | Bej et al., 1991 | ||
| Pseudomonas aeruginosa | oprL | F: ATGGAAATGCTGAAATTCGGC | ||
| R: CTTCTTCAGCTCGACGCGACG | 504 | De Vos et al., 1997 | ||
| Proteus mirabilis | ureR | F: GCGTTATCGTGTTGCTGTG | ||
| R: CATGTTTGCTGTGGTGTAGC | 225 | Wenner et al., 2002 | ||
| Klebsiella pneumoniae | khe | F: ATGCGAAACGACGGTAGTTC | ||
| R: TTACTGCTGCCGTTGATGTC | 441 | Brisse et al., 2004 | ||
| Mec A | F: AAA ATC GAT GGT AAA GGT TGG C R: AGT TCT GCA GTA CCG GAT TTG C | 533 | Oliveira et al., 2002 |
Biofilm Detection Using the Microtiter Plate Method
Biofilm formation was assessed using the microtiter plate assay as described previously, with minor modifications [14]. Standardised bacterial suspensions were incubated in 96-well plates, washed to remove planktonic cells, stained with crystal violet, and quantified by measuring optical density at 590 nm. Isolates were categorised as strong, moderate, or weak biofilm producers based on OD₅₉₀ values.
Results
Thirty-three environmental samples were analysed from male and female surgical wards. Seventeen samples were collected from the male ward and sixteen from the female ward. The recovered bacterial species included Escherichia coli, Proteus mirabilis, Pseudomonas aeruginosa, Klebsiella pneumoniae, and Staphylococcus aureus (Table 1).
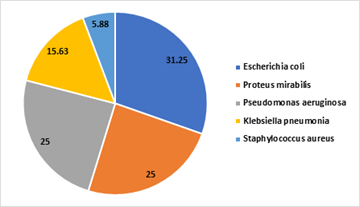
A total of 33 bacterial isolates were recovered. E. coli was the most prevalent organism (31.25%), followed by Proteus mirabilis and Pseudomonas aeruginosa (25 percentage each). Klebsiella pneumoniae accounted for 15.63 percentage, while Staphylococcus aureus was the least frequent (5.88%). Tables and drawers were the most contaminated surfaces, whereas beddings showed the lowest bacterial recovery (Figure 1; Table 2).
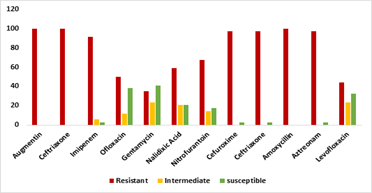
Antibiotic susceptibility testing revealed widespread multidrug resistance among the isolates. E. coli demonstrated the highest resistance levels, followed by Staphylococcus aureus, Proteus mirabilis, Pseudomonas aeruginosa, and Klebsiella pneumoniae. Overall resistance was highest to amoxicillin–clavulanate, cefotaxime, cefuroxime, and amoxicillin, while relatively lower resistance was observed to imipenem, ofloxacin, and gentamicin (Figure 2).
Carbapenem resistance was common among the isolates, with meropenem showing the highest resistance rate (80percentage), followed by ertapenem (68%) and imipenem (36percentage). Tables and bed surfaces harboured the highest proportion of carbapenem-resistant isolates (Table 3).
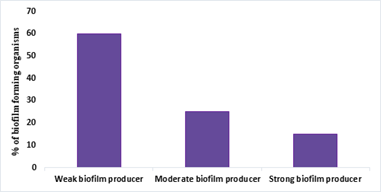
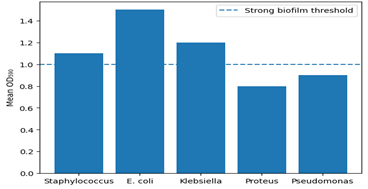
All multidrug-resistant isolates assessed demonstrated biofilm-forming ability. Sixty percent were classified as strong biofilm producers, 25% as moderate, and 15% as weak. E. coli, Klebsiella pneumoniae, and Staphylococcus species exhibited particularly strong biofilm-forming capacity (Figure 3a and 3b).
Table 1: Distribution of Environmental Samples and Bacterial Isolates from Male and Female Wards
| Ward | Surface | No. of Samples | Bacterial Isolates Recovered | Total Isolates per Ward |
| Male | Chairs | 4 | Pseudomonas aeruginosa, Escherichia coli, Staphylococcus aureus | 5 |
| Male | Tables | 5 | Klebsiella pneumoniae, Proteus mirabilis, Staphylococcus aureus | 5 |
| Male | Drawers | 5 | Pseudomonas aeruginosa, Escherichia coli | 2 |
| Male | Iron surfaces | 3 | Proteus mirabilis, Staphylococcus aureus | 2 |
| Male | Beddings | 0 | – | 0 |
| Female | Chairs | 2 | Escherichia coli, Staphylococcus aureus | 2 |
| Female | Tables | 7 | Pseudomonas aeruginosa, Klebsiella pneumoniae, Proteus mirabilis | 3 |
| Female | Drawers | 3 | Escherichia coli, Staphylococcus aureus | 2 |
| Female | Iron surfaces | 3 | Pseudomonas aeruginosa, Staphylococcus aureus | 2 |
| Female | Beddings | 1 | Klebsiella pneumoniae | 1 |
Note: Total isolates per ward indicate the cumulative number of organisms recovered from all surfaces within each ward.
Figure 1: Distribution of Isolates recovered from the Study
Table 2: Percentage Distribution of Bacterial Isolates in Male and Female Surgical Ward Environments
| Chairs | Tables | Irons | Drawers | Beddings | Total | |
| Escherichia coli | 2(33.3%) | 2(16.6%) | 1(16.6%) | 5(60.5%) | 0(0%) | 10(30.30%) |
| Proteus mirabilis | 1(16.6%) | 3(25%) | 2(33.3%) | 2(25%) | 0(0%) | 8(24%) |
| Pseudomonas aeruginosa | 2(33.3%) | 4(33.3%) | 1(16.6%) | 1(12.5%) | 0(0%) | 8(24%) |
| Klebsiella pneumonia | 1(16.6%) | 2(16.6%) | 1(16.6%) | 0(0%) | 1(100%) | 5(15.15%) |
| Staphylococcus aureus | 0(0%) | 1(8.3%) | 1(16.6%) | 0(0%) | 0(0%) | 2(6.06%) |
| Total | 6 | 12 | 6 | 8 | 1 | 33(100%) |
Figure 2: Antibiotic Susceptibility Patterns of the Isolates to the tested Antibiotics
Table 3: Percentage Prevalence of Carbapenem Resistance
| Antibiotics | Drawers n = 7 | Table n = 9 | Chairs n = 5 | Iron n = 3 | Bed n = 1 | TOTAL |
| IMP | 0(0%) | 5(13.8%) | 2(10%) | 1(8.3%) | 1(25%) | 9(36%) |
| ETP | 4(14.2%) | 7(19.4%) | 3(15%) | 2(16.6%) | 1(25%) | 17(68%) |
| MEM | 5(17.8%) | 8(22.2%) | 4(20%) | 2(16.6%) | 1(25%) | 20(80%) |
Keys: Imipenem 10μg (IMP), Meropenem 10μg (MEM), Ertapenem 10μg (ETP)
Figure 3a: Distribution of biofilm-producing capacity among bacterial isolates.
Bar chart showing the percentage of isolates classified as strong, moderate, and weak biofilm producers based on OD₅₉₀ measurements.
Figure 3b: Biofilm formation among multidrug-resistant (MDR) bacterial isolates.
Bar chart showing mean OD₅₉₀ values for MDR isolates by species. The dashed horizontal line represents the OD₅₉₀ threshold (1.0) for strong biofilm production.
Discussion
Nosocomial infections continue to pose a significant public health challenge, particularly in low- and middle-income countries, where infection control infrastructure and antimicrobial stewardship are often inadequate [15, 16]. The present study highlights the hospital environment as an important reservoir of multidrug-resistant and biofilm-forming pathogens in surgical wards.
The recovery of clinically significant bacteria from high-touch surfaces, such as tables and drawers, portrays their role in pathogen transmission within healthcare settings. Similar findings have been reported in recent studies demonstrating that contaminated environmental surfaces significantly contribute to cross-infection in hospitals [17,18]. The predominance of P. aeruginosa and S. aureus reflects their well-documented ability to survive under harsh environmental conditions and resist commonly used disinfectants [12,19].
The high prevalence of multidrug resistance, particularly to β-lactam antibiotics, is consistent with previous reports from Nigerian healthcare facilities and reflects prolonged selective pressure arising from inappropriate antibiotic use [20]. The detection of carbapenem-resistant isolates is especially concerning, as carbapenems are often considered last-resort antibiotics. Resistance rates observed in this study exceed those reported in many high-income countries [21], highlighting the urgent need for strengthened surveillance and infection control measures [22,23].
Biofilm formation represents a critical virulence factor that contributes to both antimicrobial resistance and environmental persistence of nosocomial pathogens. Biofilm development begins with bacterial adhesion to surfaces, facilitated by the adsorption of organic molecules that modify surface physicochemical properties and promote microbial attachment [24]. Once established, biofilms confer substantial protection against antibiotics, disinfectants and host immune responses, thereby complicating infection management [25,26. In the present study, all isolates demonstrated biofilm-forming capacity, with E. coli and K. pneumoniae emerging as the strongest producers. Overall, 60% of isolates were classified as strong biofilm producers, 25% as moderate, and 15% as weak, indicating a high potential for persistence on hospital surfaces. These findings are comparable with recent studies reporting high rates of biofilm formation among clinical and environmental isolates, including 88% reported by Velmurugan and Krishnasamy [27] and 100% in clinical specimens reported by Ghasemian et al. [28]. A strong association between biofilm production and multidrug resistance was observed, as most strong biofilm-forming isolates exhibited resistance to three or more classes of antibiotics. Similar correlations have been widely reported [29-32], reinforcing the role of biofilms in the persistence and dissemination of MDR pathogens. However, contrasting observations by Qi et al. [33] suggest that robust biofilm formation may also occur in non-MDR isolates, indicating that the relationship between resistance and biofilm formation is complex and multifactorial. These findings highlight the need for molecular studies to elucidate the genetic and regulatory mechanisms underlying biofilm-associated resistance.
This study provides important local epidemiological data that can inform infection prevention and control strategies at Tertiary health care facilities. The low levels of antimicrobial susceptibility observed underscore the importance of continuous resistance surveillance and the use of local epidemiological data to guide empirical therapy. Furthermore, targeted educational interventions for healthcare workers on the risks associated with biofilm-forming pathogens and strict adherence to hygiene and environmental decontamination protocols are essential for reducing the burden of HAIs [34].
Conclusion
This study demonstrates the presence of clinically important, multidrug-resistant, and biofilm-forming bacteria on environmental surfaces within male and female surgical wards of a tertiary healthcare facility. The findings portray the urgent need for improved infection prevention and control practices, routine environmental surveillance, and robust antimicrobial stewardship programmes to mitigate the burden of healthcare-associated infections.
Conclusion
Availability of Data and Material
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request
Conflict of interests
The authors declare that they have no conflict of interest in this research and publication.
Funding
This research did not receive any grant from funding sources in the public, commercial, or not-for-profit sectors. The authors personally funded it.
References
- Ijioma, C. E., Abali, I. O., Ekeleme, N. C., Orji, O. J., Uwalaka, I. W., et al. (2023). Evaluation of healthcare workers’ knowledge, attitude, and practices of aseptic techniques in primary health care centres in Edo State, Nigeria. Asian Journal of Medical Principles and Clinical Practice, 6(2):119–134.
Publisher | Google Scholor - Etemad, M., Khani, Y., Hashemi-Nazari, S. S., Izadi, N., Eshrati, B., & Mehrabi, Y. (2021). Survival rate in patients with ICU-acquired infections and its related factors in Iran’s hospitals. BMC Public Health, 21(1):787.
Publisher | Google Scholor - Nwankwo, E. O. (2018). Epidemiology of healthcare-associated infections in Nigeria: A systematic review and meta-analysis. BMJ Open, 8(6):e020431.
Publisher | Google Scholor - Zhang, Y., Wang, J., Li, X., Chen, H., & Liu, M. (2022). Practical manual of nosocomial infection prevention and control of infectious diseases. Yunnan Publishing House.
Publisher | Google Scholor - Gozel, M. G., Hekimoglu, C. H., Gozel, E. Y., Batir, E., McLaws, M. L., & Mese, E. A. (2021). National infection control program in Turkey: The healthcare-associated infection rate experiences over 10 years. American Journal of Infection Control, 49(7):885–892.
Publisher | Google Scholor - Liu, H. Y., Prentice, E. L., & Webber, M. A. (2024). Mechanisms of antimicrobial resistance in biofilms. npj Antimicrobials Resistance, 2:27.
Publisher | Google Scholor - Airaodion, A. I., Megwas, A. U., Ekenjoku, J. A., Ngwogu, K. O., & Ngwogu, A. C. (2020). Therapeutic effect of ethanolic and aqueous extracts of Ocimum gratissimum leaves against gram-positive and gram-negative bacteria. International Journal of Medical Science and Health Research, 4(2):25–31.
Publisher | Google Scholor - Airaodion, A. I., Ngwogu, A. C., Ngwogu, K. O., Ekenjoku, J. A., & Megwas, A. U. (2020). Pharmacotherapeutic activity of Allium sativum (garlic) bulb against gram-positive and gram-negative bacteria. Asian Journal of Research in Infectious Diseases, 3(3):22–27.
Publisher | Google Scholor - Karygianni, L., Ren, Z., Koo, H., & Thurnheer, T. (2020). Biofilm matrixome: Extracellular components in structured microbial communities. Trends in Microbiology, 28:668–681.
Publisher | Google Scholor - Loveday, H. P., Wilson, J. A., Pratt, R. J., Golsorkhi, M., Tingle, A., Bak, A., Browne, J., Prieto, J., & Wilcox, M. (2014). epic3: National evidence-based guidelines for preventing healthcare-associated infections in NHS hospitals in England. Journal of Hospital Infection, 86(Suppl. 1):S1–S70.
Publisher | Google Scholor - Haque, M., McKimm, J., Sartelli, M., Dhingra, S., Labricciosa, F. M., Islam, S., Jahan, D., Nusrat, T., Chowdhury, T. S., Coccolini, F., Iskandar, K., Catena, F., & Charan, J. (2020). Strategies to prevent healthcare-associated infections: A narrative overview. Risk Management and Healthcare Policy, 13:1765–1780.
Publisher | Google Scholor - Cheesebrough, F. A. (2012). Antimicrobial susceptibility survey of Pseudomonas aeruginosa strains isolated from clinical sources. Journal of the National Medical Association, 96(8):65–69.
Publisher | Google Scholor - Clinical and Laboratory Standards Institute. (2021). Performance standards for antimicrobial susceptibility testing (31st ed., CLSI Supplement M100). CLSI.
Publisher | Google Scholor - Christensen, O., Atlas, R. M., & Snyder, J. W. (2020). Handbook of media for clinical microbiology (2nd ed.,) 278–339:455.
Publisher | Google Scholor - Akhtar Danesh, L., Saiedi Nejad, Z., Sarmadian, H., Fooladvand, S., van Belkum, A., & Ghaznavi-Rad, E. (2019). Elimination of Staphylococcus aureus nasal carriage in intensive care patients lowers infection rates. European Journal of Clinical Microbiology & Infectious Diseases, 38(1):87–92.
Publisher | Google Scholor - World Health Organization. (2023). Global report on infection prevention and control. WHO Press.
Publisher | Google Scholor - Otter, J. A., Donskey, C., Yezli, S., Douthwaite, S., Goldenberg, S. D., & Weber, D. J. (2022). Transmission of SARS-CoV-2 and other pathogens via the hospital environment: A systematic review. Journal of Hospital Infection, 120:1–12.
Publisher | Google Scholor - Airaodion, A. I., Oluba, S. O., Adedeji, A. A., Emaleku, S. A., Osunmuyiwa, O. J., Megwas, A. U., & Ayita, E. B. (2021). Velvet tamarind (Dialium guineense Wild) fruit is potent against gram-positive and gram-negative bacteria. Asian Journal of Medical Principles and Clinical Practice, 4(2):230–235.
Publisher | Google Scholor - Dancer, S. J. (2021). Controlling hospital-acquired infection: Focus on the role of the environment and new technologies for decontamination. Clinical Microbiology Reviews, 34(3):e00020-19.
Publisher | Google Scholor - Aibinu, I., Odugbemi, T., Koenig, W., & Ghebremedhin, B. (2012). Extended-spectrum β-lactamase enzymes in clinical isolates of Enterobacter species from Lagos, Nigeria. Journal of Clinical Microbiology, 50(2):559–564.
Publisher | Google Scholor - Sader, H. S., Carvalhaes, C. G., Streit, J. M., Doyle, T. B., & Castanheira, M. (2021). Antimicrobial activity of ceftazidime-avibactam, ceftolozane-tazobactam, and comparators against Pseudomonas aeruginosa and Klebsiella pneumoniae isolates from United States medical centers, 2016–2018. Microbial Drug Resistance, 27(3):342–349.
Publisher | Google Scholor - Centers for Disease Control and Prevention. (2022). Antibiotic resistance threats in the United States, 2022. U.S. Department of Health and Human Services.
Publisher | Google Scholor - World Health Organization. (2024). Global antimicrobial resistance and use surveillance system (GLASS) report. WHO Press.
Publisher | Google Scholor - Murga, R., Miller, J. M., & Donlan, R. M. (2001). Biofilm formation by gram-negative bacteria on central venous catheter connectors: Effect of conditioning films in a laboratory model. Journal of Clinical Microbiology, 39(6):2294–2297.
Publisher | Google Scholor - Donlan, R. M., & Costerton, J. W. (2001). Biofilms: Survival mechanisms of clinically relevant microorganisms. Clinical Microbiology Reviews, 15(2):167–193.
Publisher | Google Scholor - Flemming, H.-C., van Hullebusch, E. D., Neu, T. R., Nielsen, P. H., Seviour, T., Stoodley, P., Wingender, J., Wuertz, S., & Szewzyk, U. (2023). The biofilm matrix: Multitasking in a shared space. Nature Reviews Microbiology, 21(2):70–86.
Publisher | Google Scholor - Velmurugan, D., & Krishnasamy, S. (2024). Biofilm formation and antimicrobial resistance patterns among clinical bacterial isolates. Journal of Infection and Public Health, 17(1):45–53.
Publisher | Google Scholor - Ghasemian, A., Shafiei, M., & Kafil, H. S. (2023). Biofilm formation in clinical isolates: A major challenge in antimicrobial therapy. Infection and Drug Resistance, 16:301–312.
Publisher | Google Scholor - Cristina, M. L., Sartini, M., Schinca, E., Ottria, G., Casini, B., & Spagnolo, A. M. (2021). Evaluation of multidrug-resistant Pseudomonas aeruginosa in healthcare facility water systems. Antibiotics, 10:1500.
Publisher | Google Scholor - Gajdács, M., Baráth, Z., Kárpáti, K., Szabó, D., Usai, D., Zanetti, S., & Donadu, M. G. (2021). No correlation between biofilm formation, virulence factors, and antibiotic resistance in Pseudomonas aeruginosa: Results from a laboratory-based in vitro study. Antibiotics, 10(9):1134.
Publisher | Google Scholor - Hu, Y., Gao, G. F., & Zhu, B. (2021). The antibiotic resistome: Gene flow in environments, animals, and human pathogens. Nature Reviews Microbiology, 19(3):139–154.
Publisher | Google Scholor - Okafor, C. O. O., Iroha, I. R., Ude, I. U., Onuoha, S. C., Ejikeugwu, C., Ovia, K. N., Eromonsele, B. O., Agah, V. M., Okoronkwo, C., Gabriel-Ibeh, I., Okoroafor, I., & Nwachukwu, O. B. (2022). Drug resistance profile of biofilm-forming Pseudomonas aeruginosa isolated from aquatic environments in southeastern Nigeria. Environmental Challenges, 8:100530.
Publisher | Google Scholor - Qi, L., Li, H., Zhang, C., Liang, B., Li, J., Wang, L., Du, X., Liu, X., Qiu, S., & Song, H. (2016). Relationship between antibiotic resistance, biofilm formation, and biofilm-specific resistance in Acinetobacter baumannii. Frontiers in Microbiology, 7:483.
Publisher | Google Scholor - World Health Organization. (2020). Guidelines on core components of infection prevention and control programmes. WHO Press.
Publisher | Google Scholor